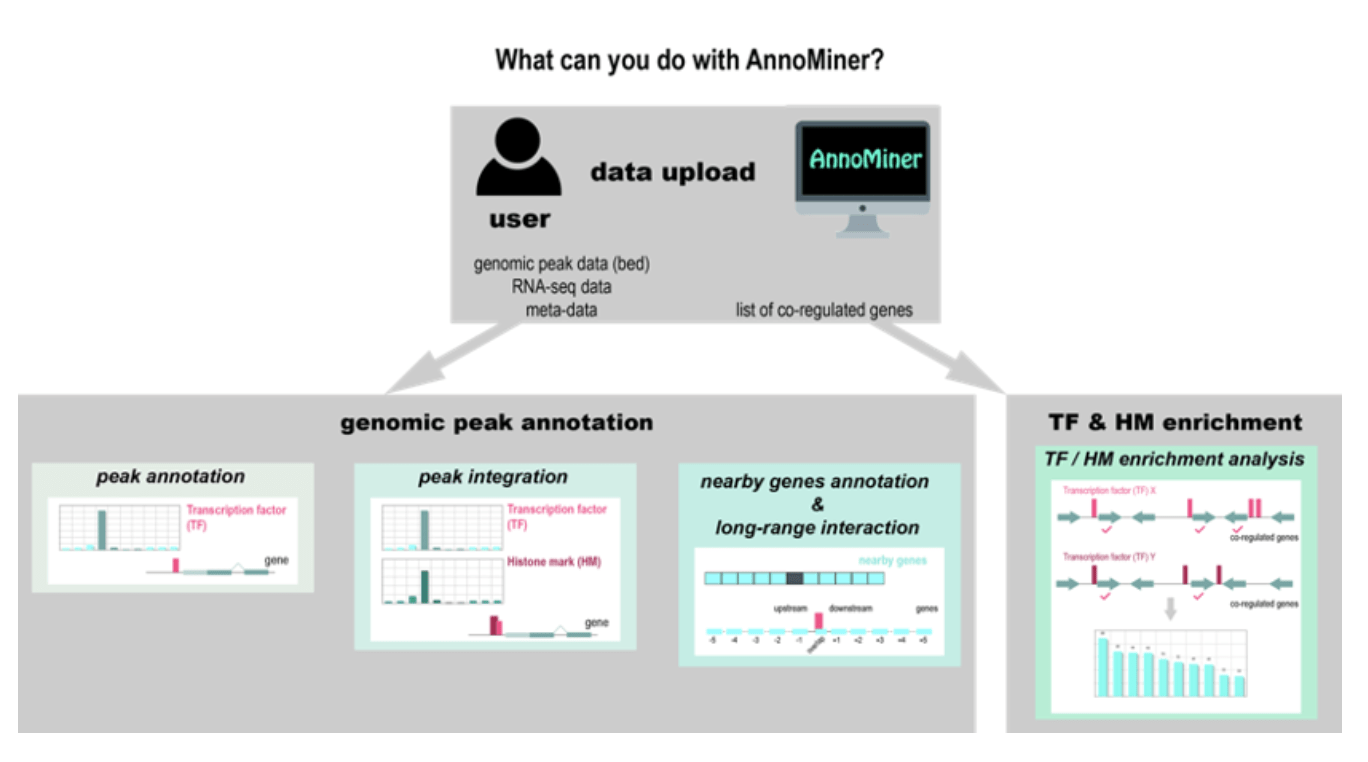
Gene regulation is a highly complex process, involving often several transcriptional regulators, sometimes over long distances on the chromosome and involving 3D chromatin structure. ChIP-seq and related techniques have helped us refine our picture of the gene regulatory process. Tools to analyze such data are often not easily accessible for wet-lab researchers, nor do they consider the complexity of this process. With AnnoMiner, we introduce a novel, user-friendly web-based tool to annotate and integrate epigenetic and transcription factor binding data. AnnoMiner gives full control to the user and you can choose, which peak – gene relationship you want to explore. Furthermore, it allows exploring genes at long distances up- or downstream from a peak. As a big bonus, you can integrate up to five genomic peak datasets with each other, and even adding information from e.g. RNA-seq on differentially expressed genes. You can also use AnnoMiner to find enriched transcription factor or histone mark peaks for a list of genes that are, for instance, co-regulated. You can find AnnoMiner here. This work is part of Fabio Marchiano’s PhD thesis (and was started already some time ago with his co-first author, Arno Meiler). It is also the product of a fruitful collaboration between the Habermann and Schnorrer teams.
Find out more: Meiler A, Marchiano F, Haering M, Weitkunat M, Schnorrer F, Habermann BH. AnnoMiner is a new web-tool to integrate epigenetics, transcription factor occupancy and transcriptomics data to predict transcriptional regulators. Sci Rep. 2021 Jul 29;11(1):15463. doi: 10.1038/s41598-021-94805-1.