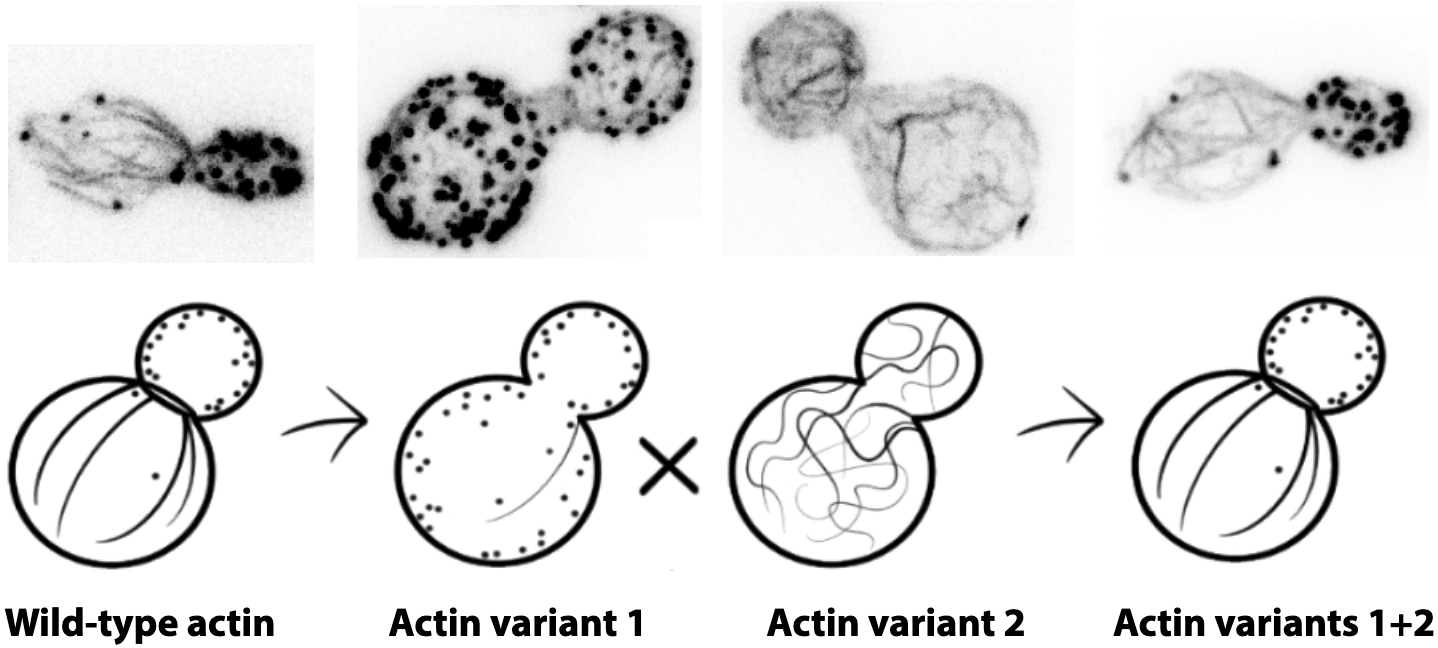
In eukaryotes, while some species assemble a complex actin cytoskeleton from a single actin ortholog (e.g. yeast), other species use a greater diversity of actin isoforms (e.g. plants). A central question of the field was how very similar actin molecules, often more than 90% identical, could nevertheless assemble into distinct networks within a common cytosplasm to perform different functions.
To answer this question, Micaela Boiero Sanders and her colleagues from the Physical and Molecular Principles of Cytoskeletal Organization team used a particular genetic model: the yeast S. cerevisiae. By replacing the only actin expressed by this organism by actins expressed in other living organisms, they showed that minor differences of actin were sufficient to globally disrupt the organization of the actin cytoskeleton by favoring the assembly of certain actin filament structures at the expense of others. Then, by expressing two complementary actins, each specialized in the assembly of distinct networks, they showed that it was possible to recover a normal organization of the cytoskeleton.
Finally, the purification of these actins and of all the proteins necessary for the assembly of the actin networks in yeast allowed to understand the molecular subtleties of these mechanisms. The authors showed that minor modifications of actin were sufficient to disrupt their interaction with actin assembly factors specific to certain actin networks, thus explaining the segregation observed in the cells.
In the end, these results allow us to propose a simple and efficient model of how life could generate new functions through the use of new actin isoforms. This model proposes that the duplication of a gene coding for actin followed by a limited number of mutations is sufficient to specialize this new actin by excluding it from certain networks. Thus, the results of this study will have a broad impact on our understanding of life.